Self-reporting of folding and aggregation by orthogonal luminophores within a single polymer chain
The folding of some polypeptides, especially enzymes, into perfectly defined three-dimensional (3D) architectures is critical to their function. The specific spatial arrangement of residues ultimately dictates function in 3D-structured proteins. Protein misfolding and aggregation are involved in the onset of many neurodegenerative age-related human disorders. To better understand diseases and design drugs, a variety of methods have been developed to investigate protein folding and to monitor the self-assembly of defective proteins into pathogenic protein aggregates. Among them, photoluminescence techniques are highly valuable due to their high sensitivity and the possibility to select, synthesize and optimize a diversity of photoluminescence probes (luminophores). These compounds allow visualizing protein folding, misfolding and aggregation in vitro and in living cells.
Trying to mimic the outstanding functionalities of proteins, the research field of folding functionalized synthetic polymers to Single-Chain NanoParticles (SCNPs) has flourished in recent years. The folded conformations of SCNPs produce robust nano-objects when stabilized by covalent bonds as surrogates of disulfide bonds in proteins. Conversely, dynamic SCNPs are obtained by folding of discrete, individual chains arising via noncovalent interactions. Current single-chain technology still lacks nature’s exquisite degree of control to generate sophisticated 3D-nanoentities. Nevertheless, some SCNPs do already mimic the outstanding properties of certain antimicrobial polypeptides as well as both the size and function of some metalloenzymes, Intrinsically Disordered Proteins (IDPs), and structural proteins.
However, as observed in the case of defective proteins, some SCNP systems are not free from multi-SCNPs´ aggregation phenomena –arising during synthesis or after long-term storage through inter-chain reactions– that negatively affect their performance, e.g., in catalysis, drug delivery and sensing applications. A visual, self-reporting method to distinguish between individual SCNPs and unbidden multi-SCNPs assemblies with the naked eye, and in the former case the degree of single-chain compaction, would be invaluable in further advancing this field of bioinspired research.
This work reports on a simple and efficient platform to identify both single-chain compaction and inter-molecular aggregation phenomena via photoluminescence based on simultaneous synthesis through Hantzsch ester formation of orthogonal luminophores within the same polymer chain (Figure, panel a). Hence, starting from non-luminescent b-ketoester-decorated chains, intra-molecular compaction is visually detected through fluorescence arising from Hantzsch fluorophores generated as intra-chain connectors during folding (Figure, panel b). Complementary, inter-molecular association is identified via Aggregation-Induced Emission (AIE) from orthogonal luminophores displaying intense photoluminescence at redshifted wavelengths after formation of multi-SCNPs assemblies (Figure, panel c). The method, which is simple and highly accurate, holds promise to perceive at a glance the presence / absence of detrimental multi-chain aggregates affecting the performance of SCNPs in catalysis, sensing, recognition, and so on.
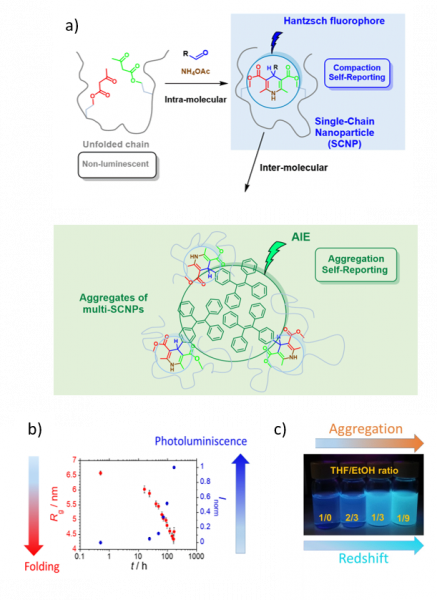
Figure: (a) Strategy for the synthesis of orthogonal luminophores based on Hantzsch ester formation within a beta-ketoester-decorated chain for self-reporting of intra-molecular compaction (folding) and inter-molecular association (aggregation). (b) Inverse relationship between single-chain size reduction, as determined by in situ SAXS kinetic measurements and photoluminescence emission observed during single-chain folding. (c) Photographs showing inter-molecular association of SCNPs in solvent (THF) / non-solvent (EtOH) mixtures at different volume ratios under 365 nm UV illumination upon increasing EtOH concentration.